Importance of the 3′-Terminal Nucleotide of the Forward Primer for Nucleoprotein Gene Detection of Viral Hemorrhagic Septicemia Virus by Conventional Reverse-Transcription PCR | SpringerLink
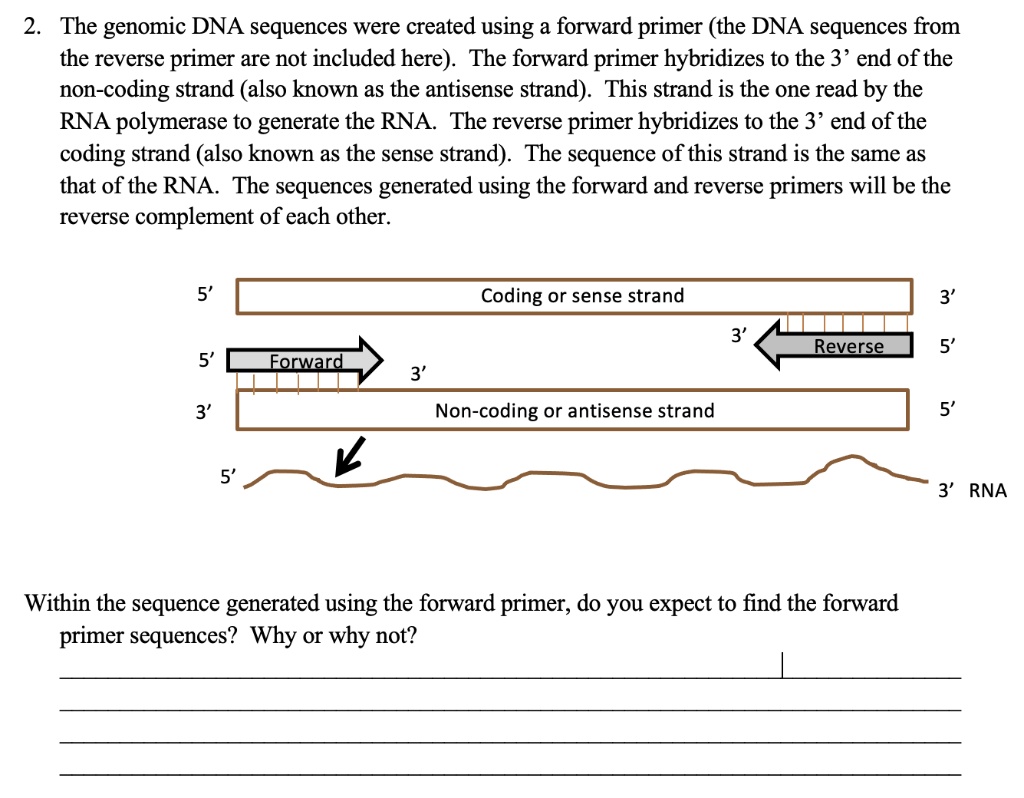
SOLVED: 2. The genomic DNA sequences were created using forward primer (the DNA sequences from the reverse primer are not included here): The forward primer hybridizes to the 3' end of the

Barcoded library preparation strategy. Forward and reverse PCR primers... | Download Scientific Diagram
Forward and reverse primers are complementary to different DNA strands. These DNA strands are complementary to each other. Which statement is right? - Quora
BatchPrimer3: A high throughput web application for PCR and sequencing primer design | BMC Bioinformatics | Full Text
A) Forward and reverse primer sequences used during PCR amplification.... | Download Scientific Diagram